Bimolecular Fluorescence Complementation(BiFC) Service
What is Bimolecular Fluorescence Complementation (BiFC)?
Bimolecular fluorescence complementation (BiFC) is a non-invasive method for studying protein-protein interactions (PPIs) in living cells. It involves splitting a fluorescent protein (e.g., YFP or GFP) into two non-fluorescent fragments, which are fused to target proteins. Upon interaction, the fragments reassemble, emitting fluorescence, which is observed via fluorescence microscopy, offering real-time evidence of PPIs.
Creative Proteomics offers a comprehensive BiFC service, providing high-quality, real-time insights into protein-protein interactions within living cells. Our service offers cutting-edge technology, flexible applications, and robust analytical tools to advance your research. Whether you're investigating cellular signaling, studying membrane proteins, or mapping protein interaction networks, BiFC is a powerful tool to help you achieve your scientific goals.
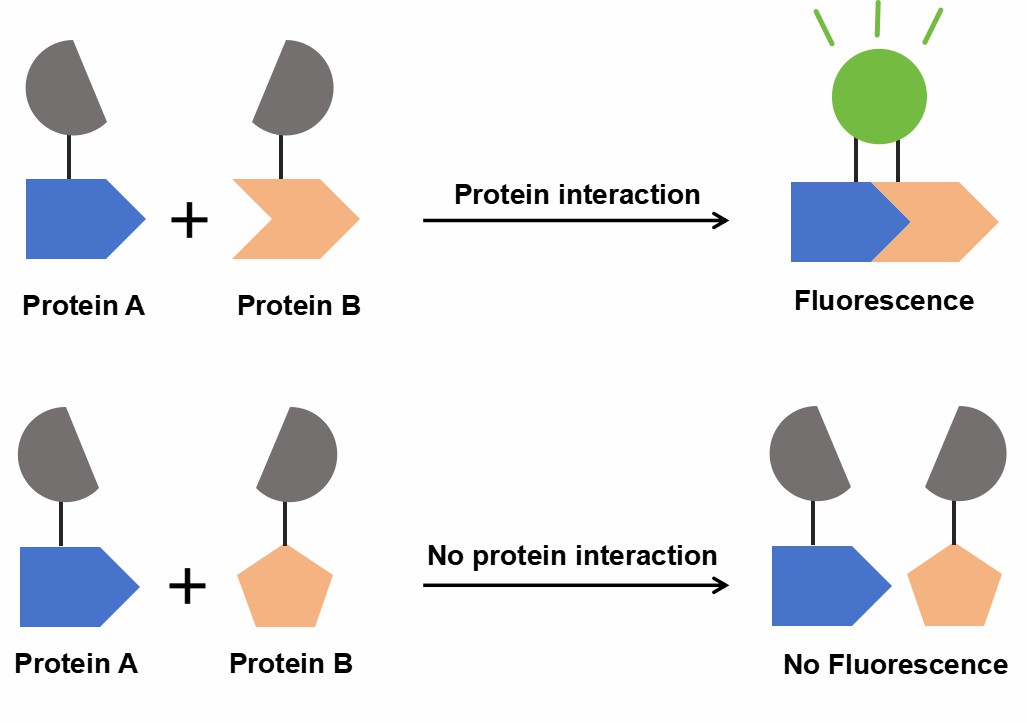
Figure 1. Schematic diagram of the principle of BiFC.
What is the difference between FRET and BiFC?
Fluorescence Resonance Energy Transfer (FRET) and BiFC are both popular techniques used for studying protein-protein interactions. However, they operate through different mechanisms and have unique applications. Below is a comparison table highlighting the key differences between FRET and BiFC:
| Feature | FRET | BiFC |
| Mechanism | Energy transfer between two fluorescent molecules | Reconstitution of a fluorescent protein from two non-fluorescent fragments |
| Type of Signal | Non-radiative energy transfer | Fluorescence emission |
| Detection | Requires donor and acceptor fluorophores | Requires fluorescence of the reformed protein |
| Interaction Detection | Can detect interactions at a distance (within ~10 nm) | Requires close proximity (direct interaction) |
| Signal Strength | Depends on the proximity of donor and acceptor fluorophores | Signal strength depends on the successful reassembly of the fluorescent protein |
| Experimental Complexity | Requires sophisticated instrumentation and calibration | Simpler setup with fewer experimental conditions |
| Sensitivity | High sensitivity for weak interactions | Suitable for both strong and weak interactions |
| Visualization | Can be used in live-cell and fixed-cell systems | Primarily used in live-cell imaging |
| Multiplexing | Can measure multiple interactions simultaneously | Typically used for one interaction at a time, though multicolor BiFC is possible |
Advantages of BiFC
Direct Visualization of Protein Interactions: BiFC allows for real-time, direct observation of protein interactions in living cells, providing clear and unambiguous results.
Non-invasive and Physiologically Relevant: Since the assay is performed in living cells, it preserves the natural cellular environment and post-translational modifications, providing a more accurate representation of protein behavior.
High Sensitivity: BiFC is highly sensitive, able to detect even weak or transient protein interactions, which are often missed by other methods.
No Need for Secondary Reagents: Unlike some techniques, BiFC does not require additional labeling agents, dyes, or antibodies, reducing potential experimental complexity.
Multiplexing Capability: The multicolor version of BiFC allows for simultaneous detection of multiple protein interactions, offering deeper insights into protein networks.
Flexible and Versatile: BiFC can be applied to a wide variety of cells and organisms, including prokaryotes, fungi, plants, and animals, making it a versatile tool for diverse biological research.
BiFC Service Workflow
Cloning: The gene of interest is cloned into an expression vector containing either the N-terminal or C-terminal fragment of a fluorescent protein.
Cell Transformation: The bait plasmid is introduced into target cells, and the cells expressing the bait protein are selected.
Library Transformation: A protein library (containing thousands of human proteins) is transformed into cells expressing the bait plasmid. These library plasmids are tagged with the complementary fragment of the fluorescent protein.
High-throughput Screening: We perform high-throughput screening using flow cytometry or a luciferase reporter system to identify protein interactions.
Fluorescence Imaging: Protein interactions are visualized through fluorescence microscopy, with fluorescence indicating successful protein interactions.
Bioinformatics Analysis: The interaction data is analyzed using our proprietary software to map the protein interaction network and identify key interactions.
Optional Validation: To further confirm interactions, additional assays like co-immunoprecipitation or cellular immunofluorescence can be performed.
Applications of BiFC
High-throughput Screening: Efficiently screen large libraries of proteins to discover novel interactions.
Protein-Protein Interaction Mapping: Map the complex networks of protein interactions within cells to understand cellular processes.
Drug Discovery: Investigate how small molecules or drugs influence protein-protein interactions, providing valuable insights for therapeutic development.
Membrane Protein Interactions: Study membrane protein interactions that are often difficult to analyze using traditional methods.
In Vivo Localization: Determine the cellular localization of interacting proteins in their natural environment.
Simple Requirement for BiFC Analysis
For Creative Proteomics' BiFC service, customers can provide the following types of samples:
| Samples types | Details |
| Cell Lines |
|
| Plasmids or Cloned Genes |
|
| Protein Samples |
|
| Small Molecules or Compounds |
|
FAQ
-
Q1: What is the sensitivity of BiFC in detecting weak protein-protein interactions?
A1: BiFC is highly sensitive and can detect even weak protein-protein interactions that might be missed by other methods. By splitting a fluorescent protein into two non-fluorescent fragments and attaching them to your proteins of interest, BiFC can reconstitute fluorescence when the proteins come into close proximity. For even weaker interactions, Creative Proteomics offers an enhanced version of the BiFC system that amplifies fluorescence signals, allowing for the detection of subtle interactions.
-
Q2: What types of proteins and cells can be used in BiFC assays?
A2: BiFC assays are versatile and can be applied to a wide range of proteins, including membrane proteins, intracellular signaling proteins, and enzymes. The method works in various cell types, from prokaryotic organisms like E. coli to eukaryotic cells (e.g., yeast, mammalian cells, plant cells). Creative Proteomics can customize assays to fit your specific experimental needs, whether you're studying a few proteins or conducting a genome-wide PPI screening.
-
Q3: Can I test multiple protein interactions at the same time?
A3: Yes, Creative Proteomics offers multicolor BiFC technology, allowing you to visualize and compare interactions between multiple proteins simultaneously in a single sample. This feature is invaluable for studying complex protein networks and understanding the interplay between different signaling pathways within a cell.
-
Case Study
Case: In Vivo Validation of Bimolecular Fluorescence Complementation (BiFC) to Investigate Aggregate Formation in Amyotrophic Lateral Sclerosis (ALS)
Abstract
This study focuses on optimizing a split-fluorescence complementation (BiFC) assay to investigate protein aggregation linked to neurodegenerative diseases, particularly amyotrophic lateral sclerosis (ALS) proteins such as TDP-43 and Fus. The research aims to provide insights into the aggregation patterns and mislocalization of these proteins in living organisms, specifically in zebrafish models. The BiFC approach offers a unique opportunity to visualize protein interactions and aggregates in vivo, providing valuable data on the mechanisms underlying ALS pathogenesis.
Methods
Fluorescence Optimization: The study used a reported optimization strategy for BiFC systems, specifically an I152L substitution in the N-terminal mVenus fragment, to reduce non-specific background fluorescence and improve signal-to-noise ratios.
mRNA Injections: Zebrafish embryos were microinjected with mRNA constructs encoding TDP-43 and Fus fused to BiFC fragments (VN155 and CC155), along with control constructs (e.g., mCerulean, mKate2). Fluorescence complementation was assessed using a plate reader and confocal microscopy.
Time Course Analysis: The temporal dynamics of the BiFC signal were observed by assessing the fluorescence intensity at multiple time points (24, 48, and 72 hpf) to track mRNA degradation and protein aggregation.
Co-Localization Studies: Co-injection with nuclear markers (e.g., Histone H2B-mCerulean3) allowed for precise localization studies of TDP-43 and Fus within zebrafish tissues.
Results
Fluorescence Optimization: The I152L substitution in the TDP-43 BiFC construct significantly reduced non-specific fluorescence, achieving a 2-fold increase in signal-to-noise ratio compared to the standard construct. This resulted in improved specificity of fluorescence complementation, making it more reliable for detecting TDP-43 interactions in vivo.
TDP-43 Localization: BiFC complementation revealed that TDP-43 primarily localizes in the nucleus but also shows some presence in the cytoplasm. The mutant TDP-43 (M337V) exhibited a higher tendency to accumulate in the cytoplasm, which is consistent with previous findings in other models of ALS.
Fus Localization and Aggregation: Wild-type Fus exhibited nuclear localization, while a mutant Fus protein (R536G) with a dysfunctional nuclear localization signal showed mislocalization to the cytoplasm. Co-expression of wild-type and mutant Fus proteins resulted in nuclear fluorescence complementation, indicating interaction between the two forms.
Temporal Dynamics of Signal: BiFC fluorescence peaked at 24-30 hours post-fertilization (hpf) and gradually declined by 72 hpf, consistent with mRNA degradation over time.
ALS Protein Interactions: The study confirmed that BiFC is a valid method for studying ALS-linked protein interactions and aggregation in vivo. Fluorescence complementation in zebrafish embryos provided insights into the formation and mislocalization of ALS-related protein aggregates, with potential applications for studying other neurodegenerative diseases.
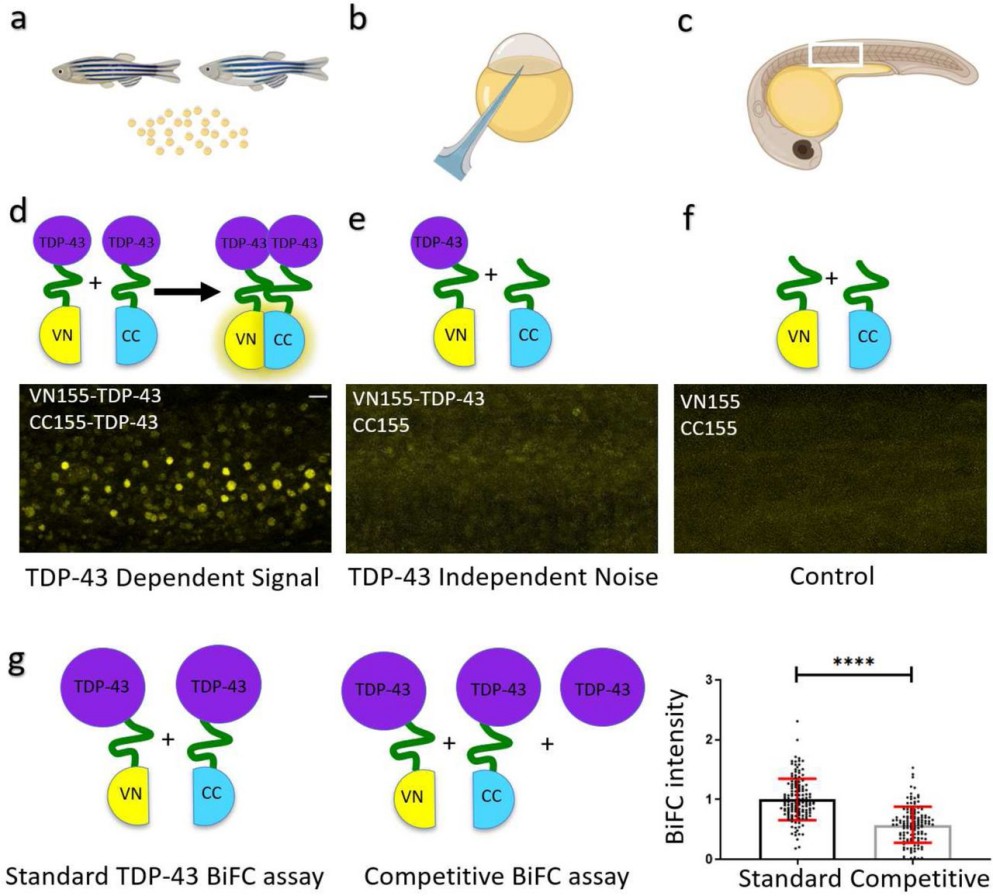
Figure 2. TDP-43 aggregation in zebrafish is specific BiFC assay to determine TDP-43 aggregation. a-c: Graphical illustration of the workflow. Male and female zebrafish are set up to collect the fertilized eggs (a). Eggs at the 1-2 cell stage are microinjected (b) with a combination of BiFC mRNA as illustrated below. Embryos are raised and BiFC complementation is visualized using a microscope (c). d: TDP-43-aggregation and respective fluorescent signal in zebrafish somites at 24hpf. e: TDP-43-independent signal (noise). f: Control (background) fluorescence at 24 hpf. Scale bar represents 20 µm. g: Comparison of BiFC intensity after standard and competitive TDP-43 BiFC injections.
Related Services
References
- Don E K, et al. In vivo validation of bimolecular fluorescence complementation (BiFC) to investigate aggregate formation in amyotrophic lateral sclerosis (ALS). Molecular Neurobiology, 2021, 58(5): 2061-2074. DOI: 10.1007/s12035-020-02238-0
- Kerppola T K. Visualization of molecular interactions using bimolecular fluorescence complementation analysis: characteristics of protein fragment complementation. Chemical Society Reviews, 2009, 38(10): 2876-2886. DOI: 10.1039/B909638H
